|
I have also tried increasing tolerance, but to no avail. 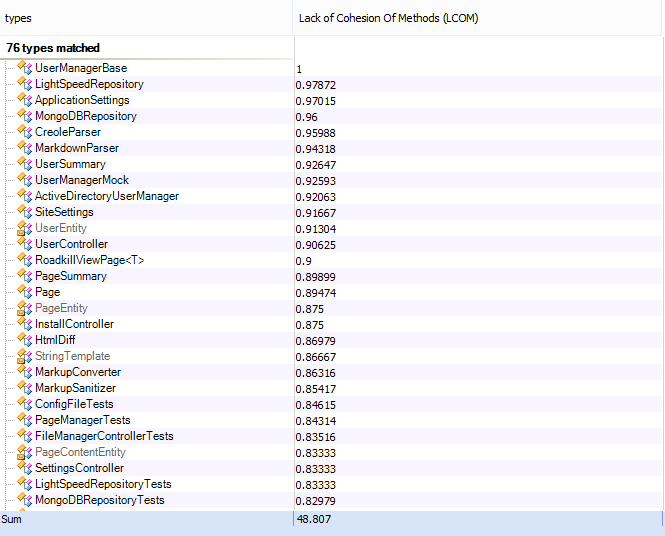
Start = StartLogistic, control=list(nlsTols=100)) Logistic_gnls = gnls(Weight ~ Asym/(1 exp(b K*Age)), data = WTS_w, I have read in a few places that increase nlsTols to 0.1 should fix the problem, but I have tried increasing it in increments of an order of magnitude up to 100, and it gives the same error. Step halving factor reduced below minimum in NLS step This is giving the error message: Error in gnls(Weight ~ Asym/(1 exp(b K * Age)), data = WTS_w, start = StartLogistic): Logistic_gnls = gnls(Weight ~ Asym/(1 exp(b K*Age)), data = WTS_gw, ggplot(walkingstick.ordered, aes(indiv, femurLength)) The stat_summary layer with geom = "linerange" draws a line from the smallest to largest measurement for each individual. Walkingstick.ordered$indiv <- rep(1:25, rep(2, 25))įinally, make the plot. walkingstick.ordered <- arrange(walkingstick, meanFemur) Finally, make a new variable indiv to label the ordered specimens. Then, order the specimens by their mean femur lengths, so that we can plot them smallest to largest. Head(ame(walkingstick)) # specimen femurLength meanFemur 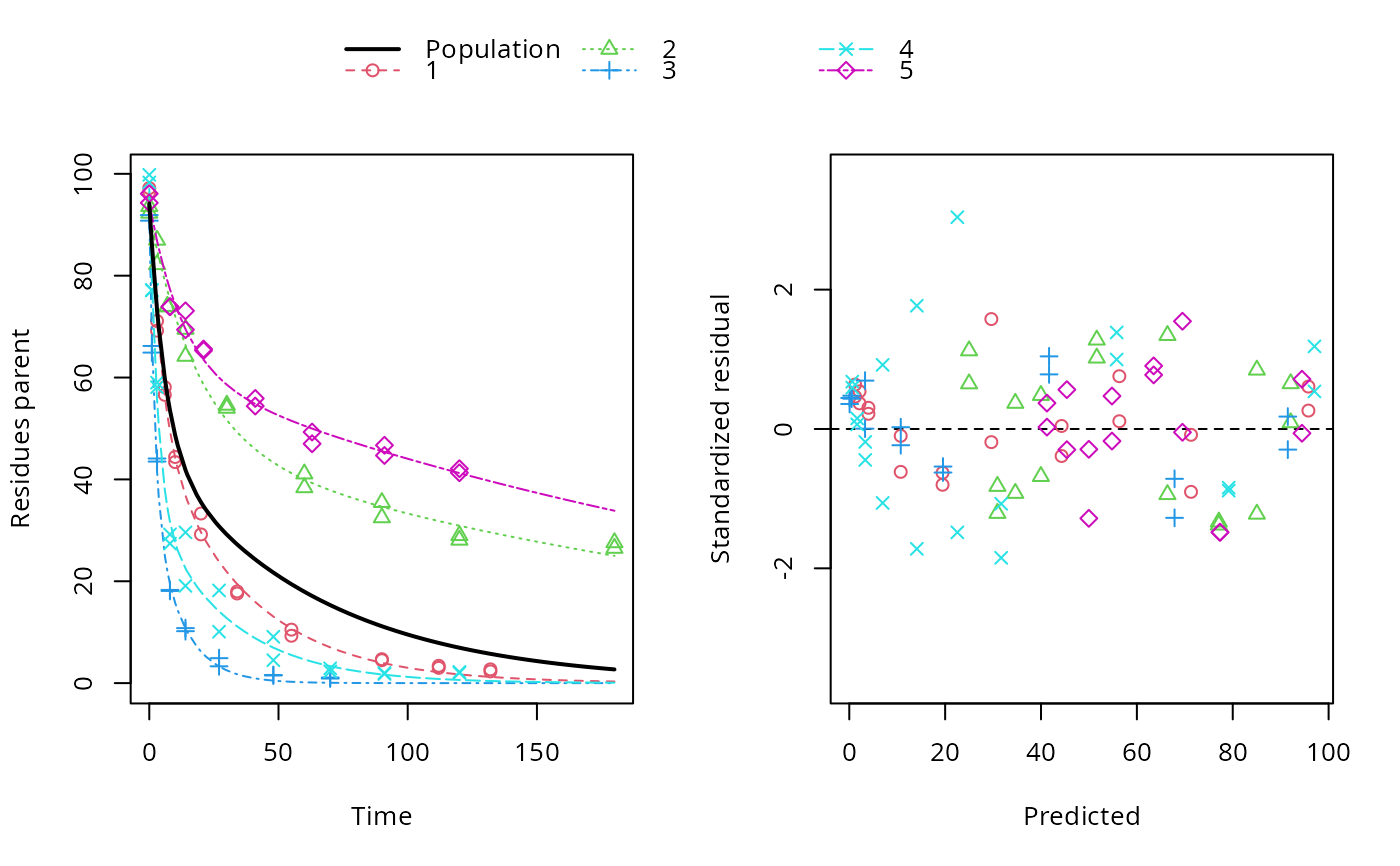 
walkingstick <- mutate(group_by(walkingstick, specimen), meanFemur = mean(femurLength)) Correlation Structure: General Formula: time id Parameter estimate (s): Correlation: 1 2 3 2 0.571 3 0.570 0.775 4 0.577 0.582 0.581 Variance function: Structure: Different standard deviations per stratum Formula: 1 time. To make the strip chart in Figure 15.6-1, we first need to order the specimens by their mean measurement.įirst, calculate the mean femur length for each specimen, as well as which of its two measurements is the smallest and which the largest. nlme is a package for fitting and comparing linear and nonlinear mixed effects models. In the summary (mod1), the following parts of the results are of interest to me that I would love to retrieve. # knee - eyes 1.216 0.376 19 3.231 0.0044 # orĬircadianPlanned # for the control vs knee comparison alone # contrast estimate SE df t.ratio p.value The planned 2-sample \(t\)-test between two means is just circadianPlanned # contrast estimate SE df t.ratio p.value Now, lme4 can easily handle very huge number of random effects (hence, number of individuals in a given study) thanks to its C part and the use of sparse matrices. # control - eyes 1.243 0.364 19 0.480 2.01Ĭonfint(circadianPlanned) # for the control vs knee comparison alone # contrast estimate SE df lower.CL upper.CL in nlme, it is possible to specify the variance-covariance matrix for the random effects (e.g. To obtain the planned 95% confidence intervals for a pairwise comparison, use circadianPlanned <- contrast(circadianPairs, method = "pairwise", adjust = "none")Ĭonfint(circadianPlanned) # contrast estimate SE df lower.CL upper.CL circadianPairs <- emmeans(circadianAnova, specs = "treatment") The only one we are interested in is that between the “control” and “knee” treatment groups, but emmeans() will calculate all pairs.įirst, apply the emmeans() command to the ANOVA object. 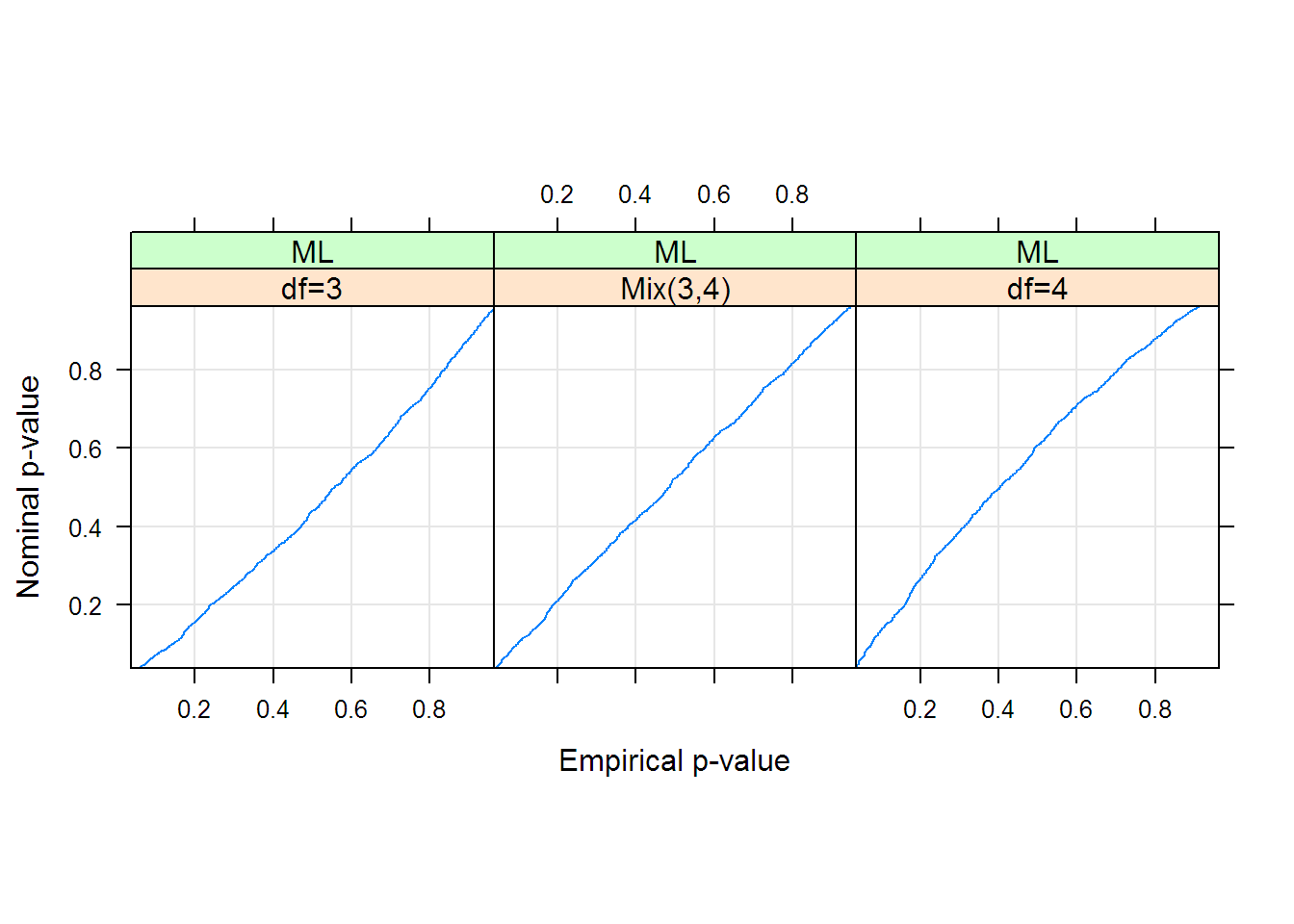
Here we’ll calculate a planned comparison.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
